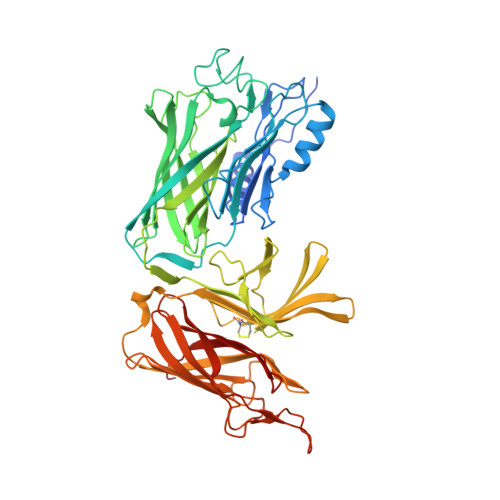
Structural Variations within the Transferrin Binding Site on Transferrin-binding Protein B, TbpB.
Calmettes, C., Yu, R.H., Silva, L.P., Curran, D., Schriemer, D.C., Schryvers, A.B., Moraes, T.F.(2011) J Biol Chem 286: 12683-12692
- PubMed: 21297163
- DOI: https://doi.org/10.1074/jbc.M110.206102
- Primary Citation of Related Structures:
3PQS, 3PQU - PubMed Abstract:
Pathogenic bacteria acquire the essential element iron through specialized uptake pathways that are necessary in the iron-limiting environments of the host. Members of the Gram-negative Neisseriaceae and Pasteurellaceae families have adapted to acquire iron from the host iron binding glycoprotein, transferrin (Tf), through a receptor complex comprised of transferring-binding protein (Tbp) A and B. Because of the critical role they play in the host, these surface-exposed proteins are invariably present in clinical isolates and thus are considered prime vaccine targets. The specific interactions between TbpB and Tf are essential and ultimately might be exploited to create a broad-spectrum vaccine. In this study, we report the structure of TbpBs from two porcine pathogens, Actinobacillus pleuropneumoniae and suis. Paradoxically, despite a common Tf target, these swine related TbpBs show substantial sequence variation in their Tf-binding site. The TbpB structures, supported by docking simulations, surface plasmon resonance and hydrogen/deuterium exchange experiments with wild-type and mutant TbpBs, explain why there are structurally conserved elements within TbpB homologs despite major sequence variation that are required for binding Tf.
Organizational Affiliation:
Department of Biochemistry, University of Toronto, Toronto, Ontario M5S 1A8, Canada.